New sequencing study links autism to FMRP, the gene disrupted in fragile X syndrome and a regulator of synaptic plasticity
Scientists reported a new connection between autism and fragile x syndrome in the latest issue of neuron. They sequenced the exomes–the parts of DNA that code proteins–of 343 families that had a single child with autism and at least one unaffected sibling. Looking at de novo mutations (ones that occurred in the sperm or egg but not found throughout the parents), they found twice as many disruptive nonsense, splice site, and frame shift mutations in the sibling with autism as in controls (59 vs. 28). Rates of disruptive mutations were higher in females (9/29) versus males (50/314) p=.07, which matches up with data from other studies on gene copy number variations (CNVs) in autism. Levels of missense mutations appeared the same in autism and control groups, but it is often hard to judge whether a missense mutation will have a significant effect on protein function with extensive testing. Most mutations came from the fathers and rates increased with the age of the father.
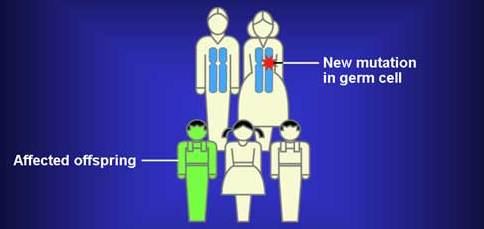
De Novo Mutations from http://www.cancer.gov/
Researchers compared the mutations they found to a previous study of gene copy number variations (CNVs) in autism. Two of the 59 disruptive mutations from the present study overlapped with the CNVs: Neurexin1 and PHF2.
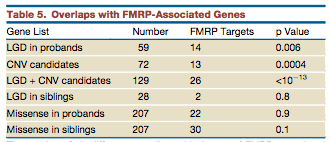
More interestingly, of the disruptive mutations in the autism group, 14/59 are part of the 842 genes regulated by FMRP (fragile X mental retardation gene product) (p=.006). Of the previously identified CNVs FMRP regulates 13/72 (p=.0004). In the disruptive mutations from controls, FMRP regulates only 2/28 a statistically insignificant amount.
When they analyzed the overall sample, and not just the de novo mutations, they also discovered an extreme scarcity of disruptive mutations in FMRP-associated genes. While they found 11.18% of synonymous variants falling within FMRP-associated genes (which would be roughly expected since they tend to be long genes), only .025% of the disruptive mutations fell into this category (P<10^-50). This strongly suggests that there is a purifying selection pressure against disruptive mutations to FMRP-associated genes.
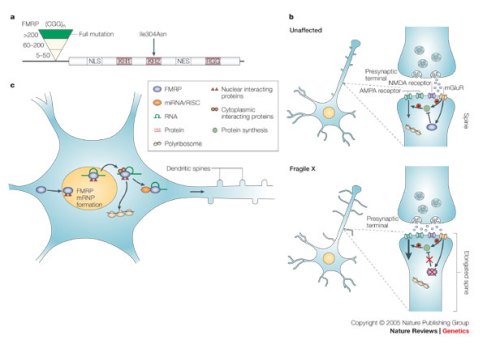
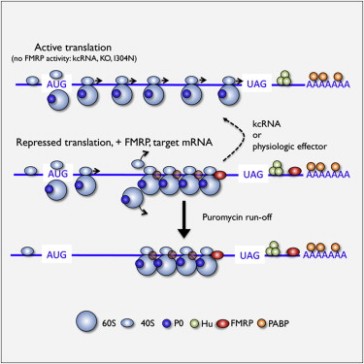
Previously autism and FMRP had been connected because 90% of children with fragile X syndrome have autistic symptoms. FMRP regulates mRNA and is thought to inhibit synaptic protein translation of transcripts involved in synaptic plasticity, which is known to be essential for learning, memory, and general brain function, so from a mechanistic standpoint it is clear that such disruptions could impact cognition in atuism. Further, other regulators of synaptic plasticity such as TSC2 (tuberous sclerosis protein 2) and synaptic scaffolding proteins have also been implicated in autism.
The researchers hope to expand the study from the 343 families to 2800 families from the Simons Simplex Collection.
——
In additions to the advances we are making in the genetics of autism (and it appears to be one of the most heritable psychiatric disorders), we are making great progress in early diagnosis. Check out the post: Using machine learning, Harvard researchers create a web-based tool to diagnose autism in minutes.
You may also be interested in other articles about the neuroscience of psychiatric disorders.
Filed under: Autism, Disease, Fragile X, Genetics, Human, Plasticity, Psychiatric | Leave a Comment
Tags: ASD, de novo mutations, exome sequencing, FMRP, genetics of psychiatric disease, rare mutations, simons simplex, synaptic plasticity


No Responses Yet to “New sequencing study links autism to FMRP, the gene disrupted in fragile X syndrome and a regulator of synaptic plasticity”